Version 3.0 of the TheoDORE wavefunction analysis package is available. Download the current version below.
New features of TheoDORE 3.0
- New user interface and documentation
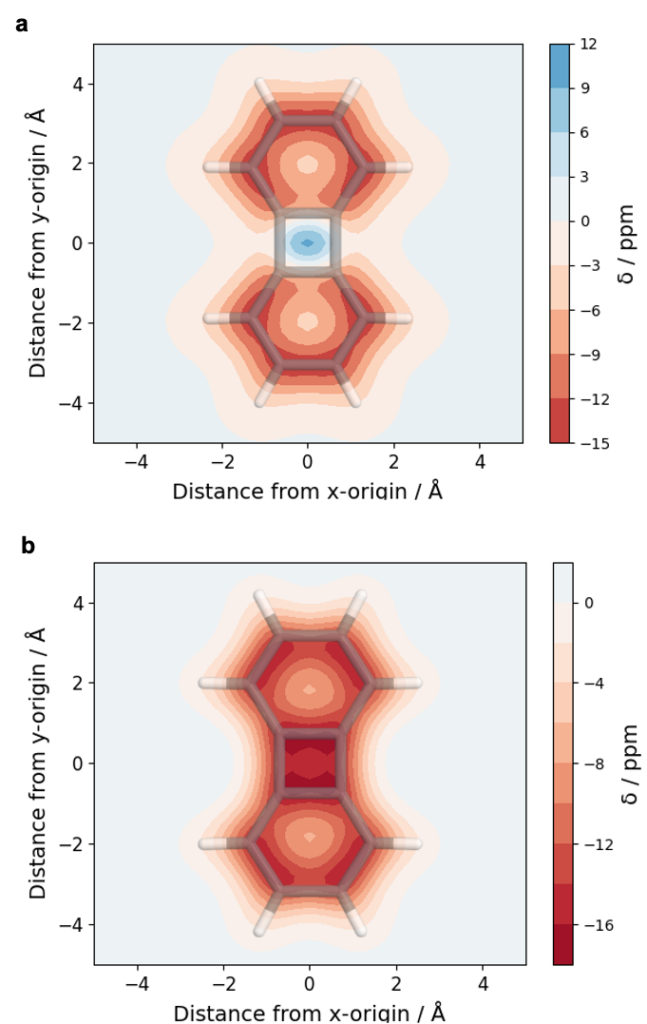
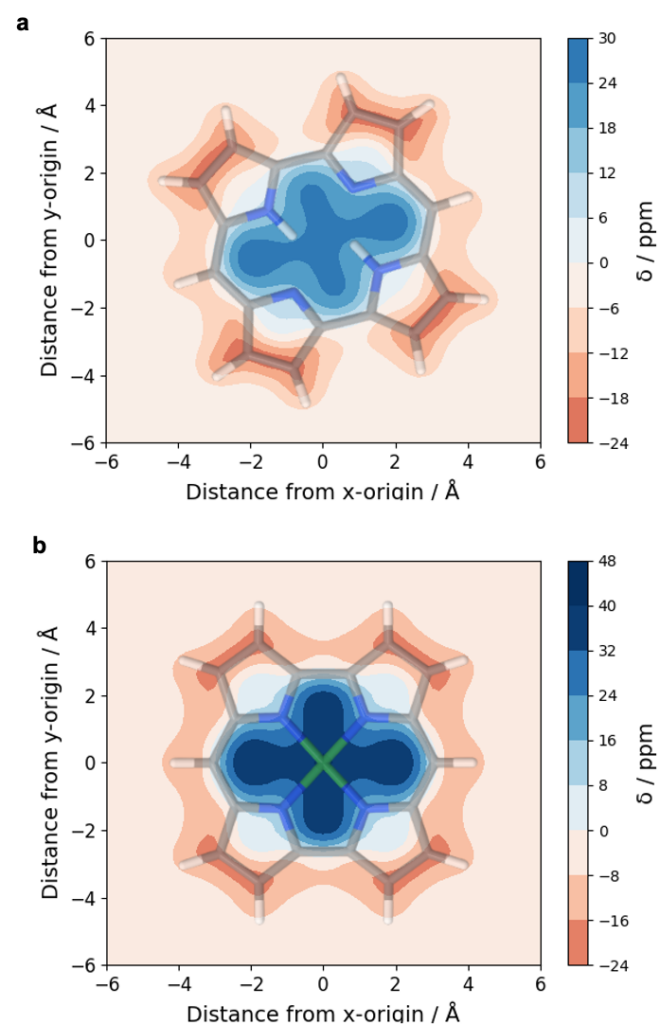
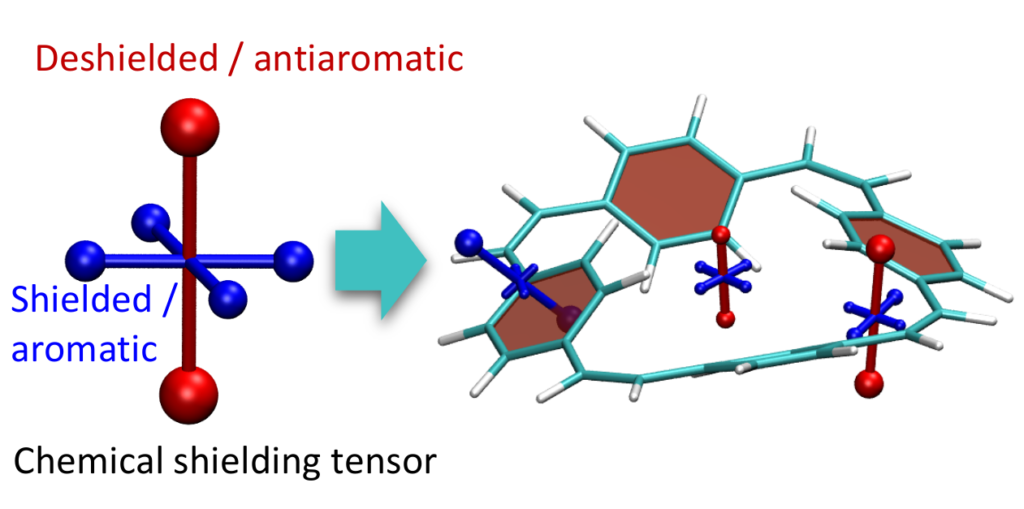
- Improvement for VIST (plot_vist)
- Improvements for natural orbital analysis (analyze_nos) including unrestricted orbitals
- LOC for ionic states (analyze_tden)
- Jmol densities (jmol_mos)
- State-to-state TDM
- Updated ADF interface
- ONETEP interface
- Excitation number, modified from [DOI: (10.1021/acs.jctc.7b00963)]
Note: TheoDORE 3 has a modified user interface. To use TheoDORE call
theodore theoinp
theodore analyze_tden
theodore analyze_nos
etc.
Download the newest release of the TheoDORE wavefunction analysis program – TheoDORE 3.1.1 (23 June 2023)